
by Graham Williams

|
Data Science Desktop Survival Guide
by Graham Williams |
|
|||
Regexp Pattern Matching |
| Metacharacter | Description | |
| 1 | ^ |
Matches at the start of the string |
| 2 | $ | Matches at the end of the string |
| 3 | () | Define a subexpression to be matched and retrieved later. |
| 4 | 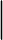 |
Matches the pattern before or pattern after |
| 5 | [ ] | Matches a single character that is contained within bracket |
| 6 | . | Matches any single character |
g/re/p is a command from the
command line tool ed to get the regular
expression and print it.
|
s <- c("hands", "data", "on", "data$cience", "handsondata$cience", "handson")
grep(pattern="^data", s, value=TRUE)
grep(pattern="on$", s, value=TRUE)
grep(pattern="(nd)..(nd)", s, value=TRUE)
In order to match a metacharacter in R we need to escap it with
|
|
grep(pattern="\\$", s, value=TRUE)
|